Sustained an infection of high-risk human papillomavirus (HR-HPVs), particularly HPV16 and HPV18,is a significant trigger of cervical cancer. E6 and E7 oncoproteins, encoded by the HPV genome, are vital for transformation and upkeep of malignant phenotypes of cervical cancer. Here, we used an rising programmable clustered repeatedly interspaced brief palindromic repeat (CRISPR)/Cas13a system to cleave HPV 16/18 E6/E7 messenger RNAs (mRNAs).
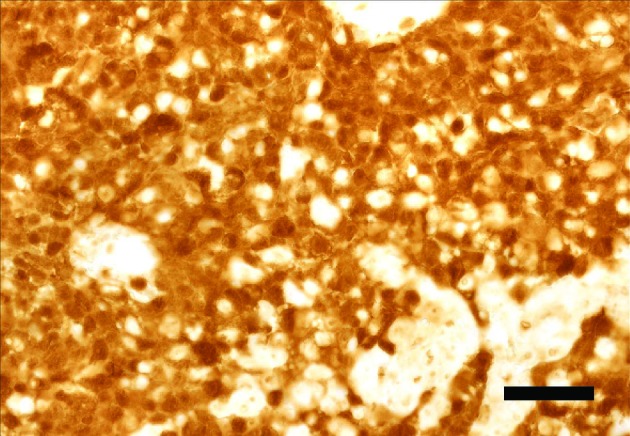
The outcomes confirmed that personalized CRISPR/Cas13a system successfully and particularly knocked down HPV 16/18 E6/E7 mRNAs, inducing growth inhibition and apoptosis in HPV16-positive SiHa and HPV18-positive HeLa Cell strains, however not in HPV-negative C33A cell line. Simultaneously, we detected downregulation of E6/E7 oncoproteins and upregulation of tumor suppressor P53 and RB proteins.
In addition, we used subcutaneous xenograft tumor growth assays to seek out that the load and quantity of tumors in the SiHa-16E6CR1 group knocked down by the CRISPR/Cas13a system have been considerably decrease than these in the SiHa-VECTOR group missing crRNA.
Our research demonstrated that focusing on HPV E6/E7 mRNAs by the CRISPR/Cas13a system could also be a candidate therapeutic technique for HPV-related cervical cancer.
Vaccination with reasonable protection eradicates oncogenic human papillomaviruses if a gender-neutral technique is utilized
Humanpapillomavirus (HPV) vaccination of ladies with very excessive (>90%) protection has the potential to eradicate oncogenic HPVs, however such excessive protection is tough to attain.
The herd impact (HE), nonetheless, relies upon each on the HPV sort and the vaccination technique.We randomized 33 Finnish communities into gender-neutral HPV16/18 vaccination, girls-only HPV16/18 vaccination, and hepatitis B-virus vaccination arms. In 2007-2010, 11,662/20,513 of 40,852/39,420 resident boys/ladies from 1992-1995 beginning cohorts consented. In 2010-14, cervicovaginal samples from vaccinated and unvaccinated ladies at age 18.5 years have been typed for HPV6/11/16/18/31/33/35/39/45/51/52/56/58/59/66/68.
Vaccine efficacy (VE) for vaccinated ladies, HE for unvaccinated ladies, and the protecting effectiveness (PE) for all ladies, have been estimated. We prolonged the community-randomized trial outcomes about vaccination technique with mathematical modeling to evaluate HPV eradication.
The HE and PE estimates in the 1995 beginning cohort for HPV18/31/33 have been important in the gender-neutral arm, and 150% and 40% stronger than in the girls-only arm. Concordantly, HPV18/31/33 eradication was predicted in adolescents/younger adults in already 20 years with 75% protection of gender-neutral vaccination.
With the 75% protection, eventual HPV16 eradication was additionally predicted, however solely with the gender-neutral technique.Gender-neutral vaccination is superior for eradication of oncogenic HPVs.
Humanpapillomavirus (HPV) an infection is a main trigger of cervical cancer. Although epidemiologic research revealed that carcinogenic danger differs in line with HPV genotypes, the expression patterns of HPV-derived transcripts and their dependence on HPV genotypes haven’t but been totally elucidated.
In this research, 382 sufferers with irregular cervical cytology have been enrolled to evaluate the associations between HPV-derived transcripts and cervical intraepithelial neoplasia (CIN) grades and/or HPV genotypes. Specifically, 4 HPV-derived transcripts, particularly, oncogenes E6 and E6*, E1^E4, and viral capsid protein L1 in 4 main HPV genotypes-HPV 16, 18, 52, and 58-were investigated.
The detection charge of E6/E6* elevated with CIN development, whereas there was no important change in the detection charge of E1^E4 or L1 amongst CIN grades. In addition, we discovered that L1 gene expression was HPV type-dependent.
Almost all HPV 52-positive specimens, roughly 50% of HPV 58-positive specimens, round 33% of HPV 16-positive specimens, and just one HPV18-positive specimen expressed L1.We demonstrated that HPV-derived transcripts are HPV genotype-dependent. Especially, expression patterns of L1 gene expression may mirror HPV genotype-dependent patterns of carcinogenesis.
Contamination of reagents and cross contamination throughout samples is a long-recognized concern in molecular biology laboratories.
While typically innocuous, contamination can result in inaccurate outcomes. Cantalupo et al., for instance, discovered HeLa-derived humanpapillomavirus18 (H-HPV18) in a number of of The Cancer Genome Atlas (TCGA) RNA-sequencing samples. This work motivated us to evaluate a better quantity of samples and decide the origin of potential contaminations utilizing viral sequences.
To detect viruses with excessive specificity, we developed the publicly out there workflow, VirDetect, that detects virus and laboratory vector sequences in RNA-seq samples. We utilized VirDetect to 9143 RNA-seq samples sequenced at one TCGA sequencing heart (28/33 cancer varieties) over 5 years.We confirmed that H-HPV18 was current in many samples and decided that viral transcripts from H-HPV18 considerably co-occurred with these from xenotropic mouse leukemia virus-related virus (XMRV).
Using laboratory metadata and viral transcription, we decided that the possible contaminant was a pool of cell strains referred to as the “frequent reference”, which was sequenced alongside TCGA RNA-seq samples as a management to observe high quality throughout know-how transitions (i.e. microarray to GAII to HiSeq), and to hyperlink RNA-seq to earlier technology microarrays that standardly used the “frequent reference”.
One of the cell strains in the pool was a laboratory isolate of MCF-7, which we found was contaminated with XMRV; one other constituent of the pool was possible HeLa cells.
Altogether, this means a multi-step contamination course of. First, MCF-7 was contaminated with an XMRV. Second, this contaminated cell line was added to a pool of cell strains, which contained HeLa. Finally, RNA from this pool of cell strains contaminated a number of TCGA tumor samples most-likely throughout library development. Thus, these human tumors with H-HPV or XMRV reads have been possible not contaminated with H-HPV 18 or XMRV.
